MOVIS: A multi-omics software solution for multi-modal time-series clustering, embedding, and visualizing tasks
@Article{Anžel2022,
author={An{\v{z}}el, Aleksandar
and Heider, Dominik
and Hattab, Georges},
title={MOVIS: A multi-omics software solution for multi-modal time-series clustering, embedding, and visualizing tasks},
journal={Computational and Structural Biotechnology Journal},
year={2022},
month={Jan},
day={01},
publisher={Elsevier},
volume={20},
pages={1044-1055},
issn={2001-0370},
doi={10.1016/j.csbj.2022.02.012},
url={https://doi.org/10.1016/j.csbj.2022.02.012}
}
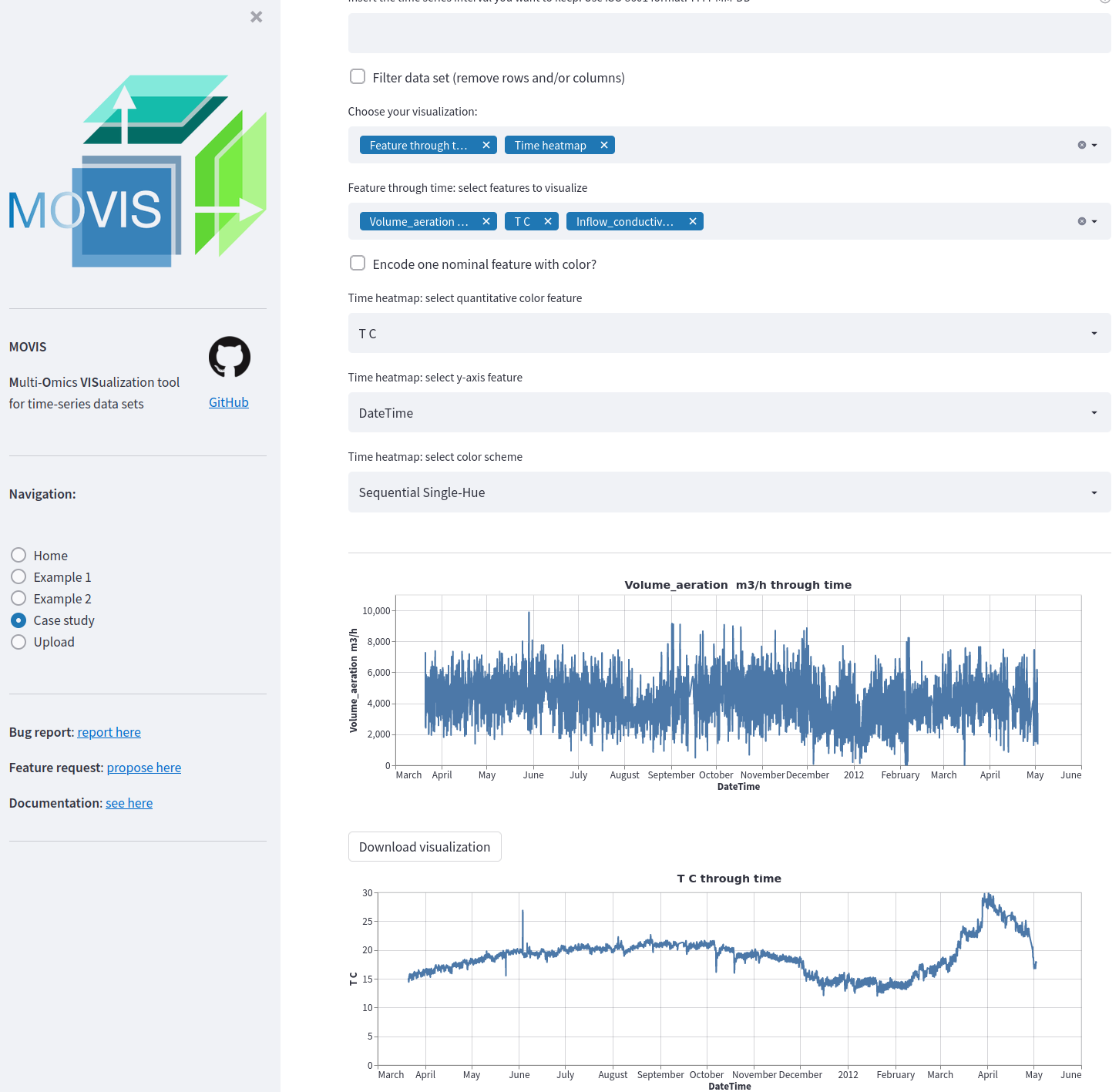
Thanks to recent advances in sequencing and computational technologies, many researchers with biological and/or medical backgrounds are now producing multiple data sets with an embedded temporal dimension. Multi-modalities enable researchers to explore and investigate different biological and physico-chemical processes with various technologies. Motivated to explore multi-omics data and time-series multi-omics specifically, the exploration process has been hindered by the separation introduced by each omics-type. To effectively explore such temporal data sets, discover anomalies, find patterns, and better understand their intricacies, expertise in computer science and bioinformatics is required. Here we present MOVIS, a modular time-series multi-omics exploration tool with a user-friendly web interface that facilitates the data exploration of such data. It brings into equal participation each time-series omic-type for analysis and visualization. As of the time of writing, two time-series multi-omics data sets have been integrated and successfully reproduced. The resulting visualizations are task-specific, reproducible, and publication-ready. MOVIS is built on open-source software and is easily extendable to accommodate different analytical tasks. An online version of MOVIS is available under https://movis.mathematik.uni-marburg.de/ and on Docker Hub (https://hub.docker.com/r/aanzel/movis).